Photosynthesis Overview
Photosynthesis involves the conversion, by various phototrophic organisms, of light energy into organic molecules, with or without the production of oxygen.
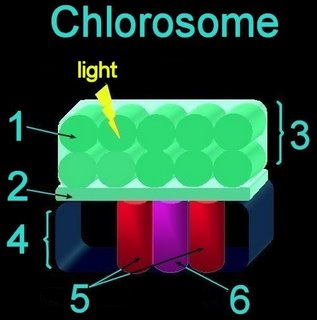
Nonoxygenic, anaerobic photosynthesis evolved^ in prokaryotes before oxygenic photosynthesis, and is found in green filamentous, green sulfur, purple sulfur, and purple nonsulfur bacteria. Photosynthesis in green sulfur bacteria such as Chlorobium tepidum takes place in the chlorosome.
Oxygenic photosynthesis apparently developed several billion years ago in an ancestor of present day Cyanobacteria. Oxygenic photosynthesis occurs in plants, which possess plastids derived from Cyanobacteria, and in photosynthetic oxygenic prokaryotes, which utilise H2O as electron donor (Cyanobacteria, prochlorophytes).
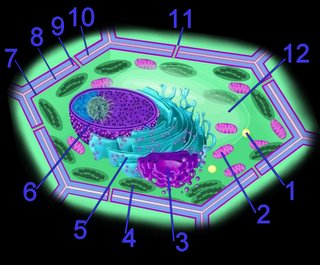
The photosystem machinery of oxygenic photosynthesis is located within the specialized internal thylakoid membrane system. The chloroplast is the site of oxygenic photosynthesis in eukaryotic cells.
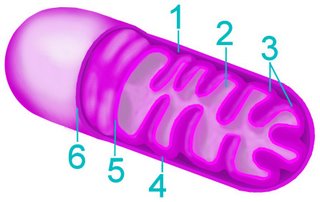
The current consensus is that chloroplasts originated from Cyanobacteria that have become endosymbionts. This is an origin analogous to the endosymbiotic (im) origin of mitochondria, which are believed derived from the "purple bacteria" (alpha-proteobacteria).
The thylakoid membrane, with its embedded membrane-bound pigments, is the structural unit of photosynthesis. Both photosynthetic prokaryotes and photosynthetic eukaryotes possess membranes with embedded photosynthetic pigments. Only eukaryotes, which have a nuclear membrane and membrane-bound organelles, have chloroplasts with an encapsulating membrane.
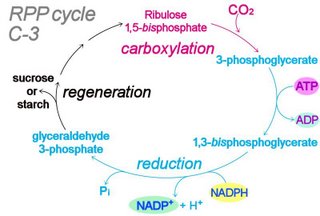
Light-dependent photosynthetic reactions employ the thylakoid membrane-embedded antenna system to harness energy delivered by a photon. The light-dependent, photophosphorylation reactions of photosystems ultimately transduce the energy of light to generate molecules of ATP and NADPH, which act as energy-transfer molecules in the light independent (“dark") reactions of the Calvin cycle. Table ~ comparison photosynthesis & respiration : Table ~ comparison plant bacterial photosynthesis : : Table ~ comparison of C-3, C-4, CAM plants :
The overall reaction of oxygenic photosynthesis is:
6 CO2 + 12 H2O → C6H12O6 + 6 O2 + 6 H2O
or,
CO2 + 2 H2O = CH2O + H2O + O2
Microbial Photosynthesis
Cyanobacteria and prochlorophytes conduct oxygenic photosynthesis. Several photosynthetic bacteria, such as the sulfur bacteria, do not generate oxygen. Nonoxygenic photosynthesis, or cyclic photophosphorylation differs from oxygenic photosynthesis in several ways other than lack of oxygen production.
Hydrogen sulfide (H2S) is utilized by the purple sulfur and green sulfur bacteria:
CO2 + 2H2S = CH2O + H2O + 2So
Nonoxygenic photosynthesis in prokaryotes, utilizing S– or So or H2 as electron donor include green filamentous, green sulfur, purple sulfur, and purple nonsulfur.
Tables Overview of Photosynthesis Comparison of C-3, C-4, CAM plants Comparison of plant and bacterial photosynthesis Structure of bacteriochlorophylls Comparison of Photosynthesis and Respiration
Diagram Z-scheme of noncyclic photophosphorylation :
HOME • • Section Photosynthesis • Calvin cycle • C-3 • C-4 • CAM • Chloroplast • Chlorosomes • Cyanobacterial cell : cyclic photophosphorylation : • Light-reactions : noncyclic photophosphorylation : • Nonoxygenic photosynthesis • Oxygenic photosynthesis • Photosynthesis Overview • Photophosphorylation • Plant cell • Timeline • • Section Pigments • Antenna and Reaction Center • Bacteriochlorophylls • Carotenoids • Chlorophylls and accessory pigments • Pigments and absorption spectra • Phycobilins Section Articles • SITE MAP • : photosynthesis ~ animations :